Details of experiments between Q9UQC2 and P31946
Note that the results of all experiments are listed, regardless of the modification states of the fragments.
Experiment series 1
Protein A protein: GAB2
Protein A fragment: 128-137
Protein A site: 14-3-3_pSer133
Protein A modification: (SEP)133
Protein A sequence: LRNVSSAGHG
Protein A construct: LRNVS(pSer)AGHG (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Measured dissociation constant (10-6M): <100
Immobilized partner concentration in holdup experiment (10-6M): 102
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 1
Measured pKd: 4
Experiment series 2
Protein A protein: GAB2
Protein A fragment: 386-395
Protein A site: 14-3-3_pThr391
Protein A modification: (TPO)391
Protein A sequence: IPRRNTLPAM
Protein A construct: IPRRN(pThr)LPAM (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.95
Holdup BI standard deviation: 0.02
Immobilized partner concentration in holdup experiment (10-6M): 102
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 8.25
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 5.28
Experiment series 3
Protein A protein: GAB2
Protein A fragment: 380-389
Protein A site: 14-3-3_pThr385
Protein A modification: (TPO)385
Protein A sequence: RSVAATIPRR
Protein A construct: RSVAA(pThr)IPRR (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.71
Holdup BI standard deviation: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 102
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 7.77
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 4.39
Experiment series 4
Protein A protein: GAB2
Protein A fragment: 380-389
Protein A site: 14-3-3_pThr385
Protein A modification: (TPO)385
Protein A sequence: RSVAATIPRR
Protein A construct: RSVAA(pThr)IPRR (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.73
Holdup BI standard deviation: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 134
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 5.82
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2022.02.17.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 4))
Number of measurements: 4
Measured pKd: 4.31
Experiment series 5
Protein A protein: GAB2
Protein A fragment: 205-214
Protein A site: 14-3-3_pSer210
Protein A modification: (SEP)210
Protein A sequence: NARSASFSQG
Protein A construct: NARSA(pSer)FSQG (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.27
Holdup BI standard deviation: 0.07
Immobilized partner concentration in holdup experiment (10-6M): 134
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 3.66
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2022.02.17.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 4))
Number of measurements: 4
Measured pKd: 3.44
Experiment series 6
Protein A protein: GAB2
Protein A fragment: 207-216
Protein A site: 14-3-3_pSer212
Protein A modification: (SEP)212
Protein A sequence: RSASFSQGTR
Protein A construct: RSASF(pSer)QGTR (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.47
Holdup BI standard deviation: 0.13
Immobilized partner concentration in holdup experiment (10-6M): 102
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 3.59
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 3.95
Experiment series 7
Protein A protein: GAB2
Protein A fragment: 205-214
Protein A site: 14-3-3_pSer210
Protein A modification: (SEP)210
Protein A sequence: NARSASFSQG
Protein A construct: NARSA(pSer)FSQG (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.33
Holdup BI standard deviation: 0.1
Immobilized partner concentration in holdup experiment (10-6M): 102
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 3.47
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 3.69
Experiment series 8
Protein A protein: GAB2
Protein A fragment: 467-476
Protein A site: 14-3-3_pSer472
Protein A modification: (SEP)472
Protein A sequence: RAGDNSQSVY
Protein A construct: RAGDN(pSer)QSVY (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: -0.13
Holdup BI standard deviation: 0.07
Immobilized partner concentration in holdup experiment (10-6M): 134
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 2.16
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2022.02.17.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 4))
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 9
Protein A protein: GAB2
Protein A fragment: 317-326
Protein A site: 14-3-3_pSer322
Protein A modification: (SEP)322
Protein A sequence: PATPLSAYQI
Protein A construct: PATPL(pSer)AYQI (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.19
Holdup BI standard deviation: 0.09
Immobilized partner concentration in holdup experiment (10-6M): 134
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 2.09
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2022.02.17.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 4))
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 10
Protein A protein: GAB2
Protein A fragment: 467-476
Protein A site: 14-3-3_pSer472
Protein A modification: (SEP)472
Protein A sequence: RAGDNSQSVY
Protein A construct: RAGDN(pSer)QSVY (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: -0.08
Holdup BI standard deviation: 0.08
Immobilized partner concentration in holdup experiment (10-6M): 102
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 1.3
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 11
Protein A protein: GAB2
Protein A fragment: 317-326
Protein A site: 14-3-3_pSer322
Protein A modification: (SEP)322
Protein A sequence: PATPLSAYQI
Protein A construct: PATPL(pSer)AYQI (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.18
Holdup BI standard deviation: 0.18
Immobilized partner concentration in holdup experiment (10-6M): 102
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 1.08
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 12
Protein A protein: GAB2
Protein A fragment: 259-268
Protein A site: 14-3-3_pSer264
Protein A modification: (SEP)264
Protein A sequence: TEFRDSTYDL
Protein A construct: TEFRD(pSer)TYDL (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: -0.05
Holdup BI standard deviation: 0.1
Immobilized partner concentration in holdup experiment (10-6M): 102
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.43
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 13
Protein A protein: GAB2
Protein A fragment: 207-216
Protein A site: 14-3-3_pSer212
Protein A modification: (SEP)212
Protein A sequence: RSASFSQGTR
Protein A construct: RSASF(pSer)QGTR (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.21
Holdup BI standard deviation: 0.42
Immobilized partner concentration in holdup experiment (10-6M): 134
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.36
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2022.02.17.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 4))
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 14
Protein A protein: GAB2
Protein A fragment: 154-163
Protein A site: 14-3-3_pSer159
Protein A modification: (SEP)159
Protein A sequence: RERKSSAPSH
Protein A construct: RERKS(pSer)APSH (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0.24
Holdup BI standard deviation: 0.42
Immobilized partner concentration in holdup experiment (10-6M): 134
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.31
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2022.02.17.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 4))
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 15
Protein A protein: GAB2
Protein A fragment: 259-268
Protein A site: 14-3-3_pSer264
Protein A modification: (SEP)264
Protein A sequence: TEFRDSTYDL
Protein A construct: TEFRD(pSer)TYDL (crude synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Average holdup BI: 0
Holdup BI standard deviation: 0.09
Immobilized partner concentration in holdup experiment (10-6M): 134
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.03
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2022.02.17.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 4))
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 16
Protein A protein: GAB2
Protein A fragment: 207-216
Protein A site: 14-3-3_pSer212
Protein A modification: (SEP)212
Protein A sequence: RSASFSQGTR
Protein A construct: biotin-ttds-RSASF(pSer)QGTR (HPLC-purified synthetic peptide)
Protein B protein: YWHAB
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3beta
Measured dissociation constant (10-6M): 116
Standard deviation of dissociation constant (10-6M): 6
Experiment method: competitive Fluorescence Polarization
Details of affinity fitting: Competitive binding equation in ProFit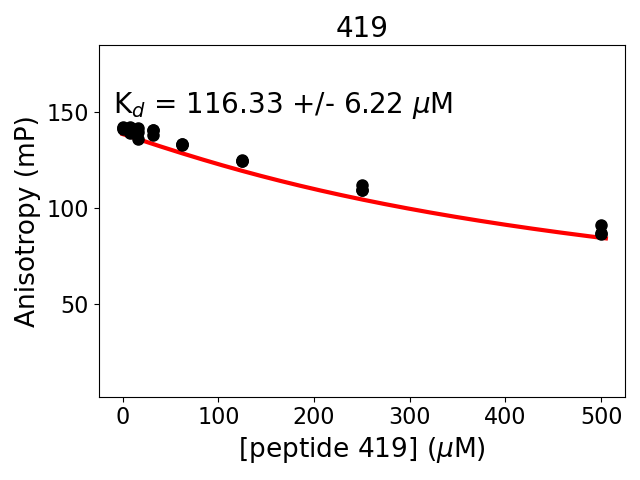
Date of measurement: 2022.02.08
Number of measurements: 3
Measured pKd: 3.94