Details of experiments between Q86UL8 and Q7Z628
Note that the results of all experiments are listed, regardless of the modification states of the fragments.
Experiment series 1
Protein A protein: MAGI2
Protein A fragment: 410-522
Protein A site: PDZ_2
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.95
Immobilized partner concentration in holdup experiment (10-6M): 35
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 5.77
Experiment series 2
Protein A protein: MAGI2
Protein A fragment: 772-865
Protein A site: PDZ_4
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.64
Immobilized partner concentration in holdup experiment (10-6M): 35
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.73
Experiment series 3
Protein A protein: MAGI2
Protein A fragment: 600-682
Protein A site: PDZ_3
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.47
Immobilized partner concentration in holdup experiment (10-6M): 35
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.42
Experiment series 4
Protein A protein: MAGI2
Protein A fragment: 1-102
Protein A site: PDZ_1
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: -0.02
Immobilized partner concentration in holdup experiment (10-6M): 35
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 5
Protein A protein: MAGI2
Protein A fragment: 915-1017
Protein A site: PDZ_5
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.06
Immobilized partner concentration in holdup experiment (10-6M): 35
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 6
Protein A protein: MAGI2
Protein A fragment: 1141-1229
Protein A site: PDZ_6
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 35
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 7
Protein A protein: MAGI2
Protein A fragment: 410-522
Protein A site: PDZ_2
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Measured dissociation constant (10-6M): 0.82
Standard deviation of dissociation constant (10-6M): 0.19
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit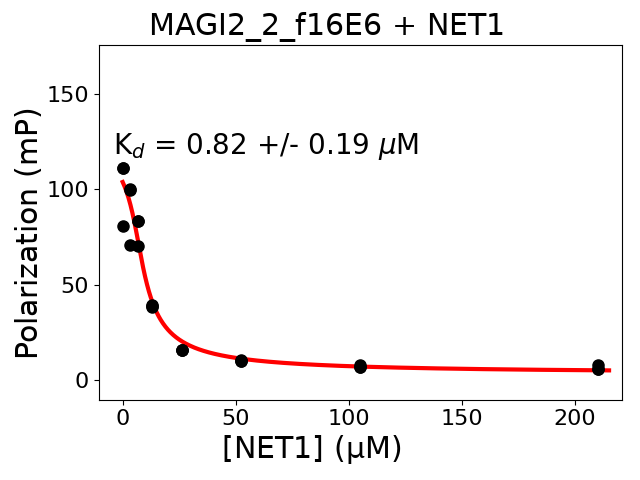
PUBMED ID of publication: 36115835
Number of measurements: 3
Measured pKd: 6.09
Experiment series 8
Protein A protein: MAGI2
Protein A fragment: 410-522
Protein A site: PDZ_2
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.83
Holdup BI standard deviation: 0.03
Normalized holdup BI: 0.91
Normalized holdup BI standard deviation: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 2
Measured pKd: 5.87
Experiment series 9
Protein A protein: MAGI2
Protein A fragment: 772-865
Protein A site: PDZ_4
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.34
Normalized holdup BI: 0.47
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.75
Experiment series 10
Protein A protein: MAGI2
Protein A fragment: 915-1017
Protein A site: PDZ_5
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.37
Normalized holdup BI: 0.37
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.56
Experiment series 11
Protein A protein: MAGI2
Protein A fragment: 600-682
Protein A site: PDZ_3
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.29
Normalized holdup BI: 0.34
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.5
Experiment series 12
Protein A protein: MAGI2
Protein A fragment: 1141-1229
Protein A site: PDZ_6
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.05
Normalized holdup BI: 0.06
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 3.54
Experiment series 13
Protein A protein: MAGI2
Protein A fragment: 410-522
Protein A site: PDZ_2
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.72
Normalized holdup BI: 0.86
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: not reported internal standards of holdup layout #5
Number of measurements: 1
Measured pKd: 5.64
Experiment series 14
Protein A protein: MAGI2
Protein A fragment: 410-522
Protein A site: PDZ_2
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.79
Normalized holdup BI: 0.87
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: not reported internal standards of holdup layout #5
Number of measurements: 1
Measured pKd: 5.66
Experiment series 15
Protein A protein: MAGI2
Protein A fragment: 600-682
Protein A site: PDZ_3
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.29
Normalized holdup BI: 0.29
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: not reported internal standards of holdup layout #5
Number of measurements: 1
Measured pKd: 4.39
Experiment series 16
Protein A protein: MAGI2
Protein A fragment: 772-865
Protein A site: PDZ_4
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.35
Normalized holdup BI: 0.42
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: not reported internal standards of holdup layout #5
Number of measurements: 1
Measured pKd: 4.65
Experiment series 17
Protein A protein: MAGI2
Protein A fragment: 915-1017
Protein A site: PDZ_5
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.41
Normalized holdup BI: 0.41
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: not reported internal standards of holdup layout #5
Number of measurements: 1
Measured pKd: 4.62
Experiment series 18
Protein A protein: MAGI2
Protein A fragment: 1141-1229
Protein A site: PDZ_6
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: biotin-ttds-SGGKRKETLV
Average holdup BI: 0.03
Normalized holdup BI: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 17.9
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: not reported internal standards of holdup layout #5
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 19
Protein A protein: MAGI2
Protein A fragment: 410-522
Protein A site: PDZ_2
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: Biotin-ado-ado-SGGKRKETLV
Average holdup BI: 0.83
Normalized holdup BI: 0.92
Immobilized partner concentration in holdup experiment (10-6M): 18
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 5.9
Experiment series 20
Protein A protein: MAGI2
Protein A fragment: 1141-1229
Protein A site: PDZ_6
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: Biotin-ado-ado-SGGKRKETLV
Average holdup BI: 0.03
Normalized holdup BI: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 18
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
PUBMED ID of publication: 36115835
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 21
Protein A protein: MAGI2
Protein A fragment: 600-682
Protein A site: PDZ_3
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: Biotin-ado-ado-SGGKRKETLV
Average holdup BI: 0.5
Normalized holdup BI: 0.52
Immobilized partner concentration in holdup experiment (10-6M): 18
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.84
Experiment series 22
Protein A protein: MAGI2
Protein A fragment: 772-865
Protein A site: PDZ_4
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: Biotin-ado-ado-SGGKRKETLV
Average holdup BI: 0.48
Normalized holdup BI: 0.64
Immobilized partner concentration in holdup experiment (10-6M): 18
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 5.06
Experiment series 23
Protein A protein: MAGI2
Protein A fragment: 915-1017
Protein A site: PDZ_5
Protein A construct: his6-MBP-TEVsite-PDZ
Protein B protein: NET1
Protein B fragment: 587-596
Protein B site: PBM
Protein B sequence: SGGKRKETLV
Protein B construct: Biotin-ado-ado-SGGKRKETLV
Average holdup BI: 0.48
Normalized holdup BI: 0.48
Immobilized partner concentration in holdup experiment (10-6M): 18
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.75