Details of experiments between Q7Z460 and Q04917
Note that the results of all experiments are listed, regardless of the modification states of the fragments.
Experiment series 1
Protein A protein: CLASP1
Protein A fragment: 277-286
Protein A site: 14-3-3_pThr282
Protein A modification: (TPO)282
Protein A sequence: RLGSSTLGSK
Protein A construct: RLGSS(pThr)LGSK (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.63
Holdup BI standard deviation: 0.05
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 6.79
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 4.11
Experiment series 2
Protein A protein: CLASP1
Protein A fragment: 1079-1088
Protein A site: 14-3-3_pThr1084
Protein A modification: (TPO)1084
Protein A sequence: RTSPLTSPTN
Protein A construct: RTSPL(pThr)SPTN (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.69
Holdup BI standard deviation: 0.08
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 6.23
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 4.24
Experiment series 3
Protein A protein: CLASP1
Protein A fragment: 682-691
Protein A site: 14-3-3_pSer687
Protein A modification: (SEP)687
Protein A sequence: KVVSQSQPGS
Protein A construct: KVVSQ(pSer)QPGS (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.71
Holdup BI standard deviation: 0.07
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 6.05
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 4.28
Experiment series 4
Protein A protein: CLASP1
Protein A fragment: 631-640
Protein A site: 14-3-3_pSer636
Protein A modification: (SEP)636
Protein A sequence: PGSYASLGRI
Protein A construct: PGSYA(pSer)LGRI (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.44
Holdup BI standard deviation: 0.04
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 5.75
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 3.77
Experiment series 5
Protein A protein: CLASP1
Protein A fragment: 589-598
Protein A site: 14-3-3_pSer594
Protein A modification: (SEP)594
Protein A sequence: VSTTGSLQRS
Protein A construct: VSTTG(pSer)LQRS (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.18
Holdup BI standard deviation: 0.04
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 4.59
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 6
Protein A protein: CLASP1
Protein A fragment: 563-572
Protein A site: 14-3-3_pSer568
Protein A modification: (SEP)568
Protein A sequence: LNRPLSAKRS
Protein A construct: LNRPL(pSer)AKRS (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: -0.47
Holdup BI standard deviation: 0.93
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.71
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 7
Protein A protein: CLASP1
Protein A fragment: 790-799
Protein A site: 14-3-3_pSer795
Protein A modification: (SEP)795
Protein A sequence: AMRVLSTSTD
Protein A construct: AMRVL(pSer)TSTD (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.04
Holdup BI standard deviation: 0.06
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.65
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 8
Protein A protein: CLASP1
Protein A fragment: 628-637
Protein A site: 14-3-3_pSer633
Protein A modification: (SEP)633
Protein A sequence: ALPPGSYASL
Protein A construct: ALPPG(pSer)YASL (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.04
Holdup BI standard deviation: 0.1
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.34
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 9
Protein A protein: CLASP1
Protein A fragment: 792-801
Protein A site: 14-3-3_pSer797
Protein A modification: (SEP)797
Protein A sequence: RVLSTSTDLE
Protein A construct: RVLST(pSer)TDLE (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.01
Holdup BI standard deviation: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.15
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 10
Protein A protein: CLASP1
Protein A fragment: 593-602
Protein A site: 14-3-3_pSer598
Protein A modification: (SEP)598
Protein A sequence: GSLQRSRSDI
Protein A construct: GSLQR(pSer)RSDI (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.01
Holdup BI standard deviation: 0.08
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.1
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 11
Protein A protein: CLASP1
Protein A fragment: 682-691
Protein A site: 14-3-3_pSer687
Protein A modification: (SEP)687
Protein A sequence: KVVSQSQPGS
Protein A construct: biotin-ttds-KVVSQ(pSer)QPGS (HPLC-purified synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Measured dissociation constant (10-6M): 42
Standard deviation of dissociation constant (10-6M): 4
Experiment method: competitive Fluorescence Polarization
Details of affinity fitting: Competitive binding equation in ProFit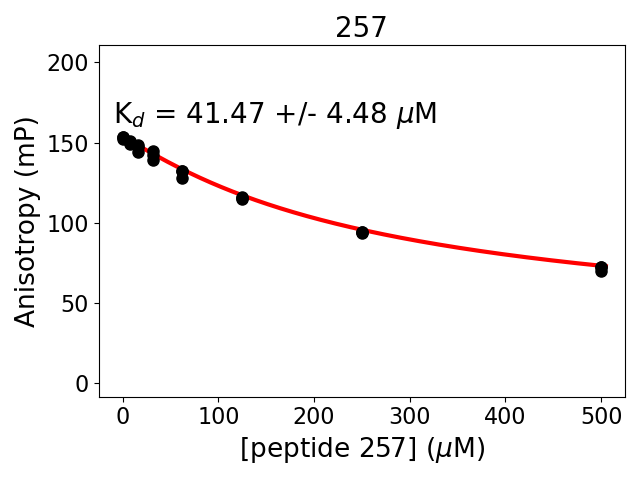
Date of measurement: 2022.01.19
Number of measurements: 3
Measured pKd: 4.38