Details of experiments between Q5VZ89 and Q04917
Note that the results of all experiments are listed, regardless of the modification states of the fragments.
Experiment series 1
Protein A protein: DENND4C
Protein A fragment: 1657-1666
Protein A site: 14-3-3_pSer1662
Protein A modification: (SEP)1662
Protein A sequence: GISTVSLPNS
Protein A construct: GISTV(pSer)LPNS (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.76
Holdup BI standard deviation: 0.06
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 7
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 4.38
Experiment series 2
Protein A protein: DENND4C
Protein A fragment: 1320-1329
Protein A site: 14-3-3_pSer1325
Protein A modification: (SEP)1325
Protein A sequence: KERSTSLSAL
Protein A construct: KERST(pSer)LSAL (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.82
Holdup BI standard deviation: 0.05
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 6.82
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 4.54
Experiment series 3
Protein A protein: DENND4C
Protein A fragment: 1273-1282
Protein A site: 14-3-3_pSer1278
Protein A modification: (SEP)1278
Protein A sequence: LERRSSLPLD
Protein A construct: LERRS(pSer)LPLD (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.54
Holdup BI standard deviation: 0.04
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 6.73
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 3.95
Experiment series 4
Protein A protein: DENND4C
Protein A fragment: 1618-1627
Protein A site: 14-3-3_pSer1623
Protein A modification: (SEP)1623
Protein A sequence: TMKISSVPNS
Protein A construct: TMKIS(pSer)VPNS (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.47
Holdup BI standard deviation: 0.04
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 6.52
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 3.83
Experiment series 5
Protein A protein: DENND4C
Protein A fragment: 1635-1644
Protein A site: 14-3-3_pSer1640
Protein A modification: (SEP)1640
Protein A sequence: LTRSHSVGGP
Protein A construct: LTRSH(pSer)VGGP (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.7
Holdup BI standard deviation: 0.12
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 4.97
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 4.25
Experiment series 6
Protein A protein: DENND4C
Protein A fragment: 1179-1188
Protein A site: 14-3-3_pSer1184
Protein A modification: (SEP)1184
Protein A sequence: RNKRSSLYGI
Protein A construct: RNKRS(pSer)LYGI (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.43
Holdup BI standard deviation: 0.1
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 4.13
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
Measured pKd: 3.76
Experiment series 7
Protein A protein: DENND4C
Protein A fragment: 1094-1103
Protein A site: 14-3-3_pSer1099
Protein A modification: (SEP)1099
Protein A sequence: IARTHSFENV
Protein A construct: IARTH(pSer)FENV (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.12
Holdup BI standard deviation: 0.05
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 2.76
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 8
Protein A protein: DENND4C
Protein A fragment: 1661-1670
Protein A site: 14-3-3_pSer1666
Protein A modification: (SEP)1666
Protein A sequence: VSLPNSLQEV
Protein A construct: VSLPN(pSer)LQEV (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.19
Holdup BI standard deviation: 0.18
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 1.07
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 9
Protein A protein: DENND4C
Protein A fragment: 1341-1350
Protein A site: 14-3-3_pSer1346
Protein A modification: (SEP)1346
Protein A sequence: VVNSLSGLKL
Protein A construct: VVNSL(pSer)GLKL (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.15
Holdup BI standard deviation: 0.25
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.53
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 10
Protein A protein: DENND4C
Protein A fragment: 960-969
Protein A site: 14-3-3_pSer965
Protein A modification: (SEP)965
Protein A sequence: IVAKHSQPSP
Protein A construct: IVAKH(pSer)QPSP (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.03
Holdup BI standard deviation: 0.1
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.23
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 11
Protein A protein: DENND4C
Protein A fragment: 984-993
Protein A site: 14-3-3_pSer989
Protein A modification: (SEP)989
Protein A sequence: IVKVPSGIFD
Protein A construct: IVKVP(pSer)GIFD (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: 0.06
Holdup BI standard deviation: 0.39
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.12
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 12
Protein A protein: DENND4C
Protein A fragment: 948-957
Protein A site: 14-3-3_pSer953
Protein A modification: (SEP)953
Protein A sequence: LIRLESIDNH
Protein A construct: LIRLE(pSer)IDNH (crude synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Average holdup BI: -0.02
Holdup BI standard deviation: 0.11
Immobilized partner concentration in holdup experiment (10-6M): 130
P value (usually -log10(P Two-sided unpaired T-test), but double check the experimental details): 0.11
Experiment method: HU Multiplex
Details of affinity fitting: hyperbolic binding equation
Date of measurement: 2021.10.13.
Experimental details: -log10(P Two-sided unpaired T-test (4 vs 5))
Number of measurements: 5
The affinity was below the detection threshold of the assay.
Experiment series 13
Protein A protein: DENND4C
Protein A fragment: 1320-1329
Protein A site: 14-3-3_pSer1325
Protein A modification: (SEP)1325
Protein A sequence: KERSTSLSAL
Protein A construct: biotin-ttds-KERST(pSer)LSAL (HPLC-purified synthetic peptide)
Protein B protein: YWHAH
Protein B fragment: 1-246
Protein B site: 14-3-3
Protein B construct: avi-his6-MBP-TEVsite-14-3-3eta
Measured dissociation constant (10-6M): 31
Standard deviation of dissociation constant (10-6M): 7
Experiment method: competitive Fluorescence Polarization
Details of affinity fitting: Competitive binding equation in ProFit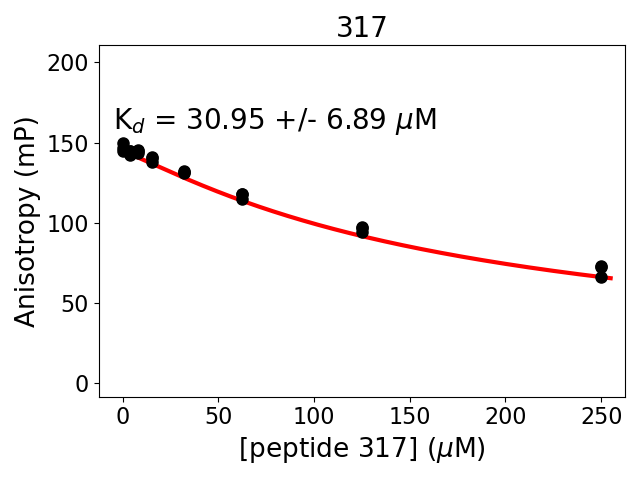
Date of measurement: 2022.01.19
Number of measurements: 3
Measured pKd: 4.51