Details of experiments between the fragments Q15418~725-735 and Q9NZN5~67-151
Note that the results of all experiments are listed, regardless of the modification states of the fragments.
Experiment series 1
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (SEP)732
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPpSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Average holdup BI: 0.14
Holdup BI standard deviation: 0.12
Immobilized partner concentration in holdup experiment (10-6M): 40
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 5
Measured pKd: 3.61
Experiment series 2
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Average holdup BI: 0.76
Holdup BI standard deviation: 0.03
Immobilized partner concentration in holdup experiment (10-6M): 31
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
PUBMED ID of publication: 36115835
Number of measurements: 8
Measured pKd: 5.06
Experiment series 3
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6M): 6.96
Standard deviation of dissociation constant (10-6M): 0.71
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit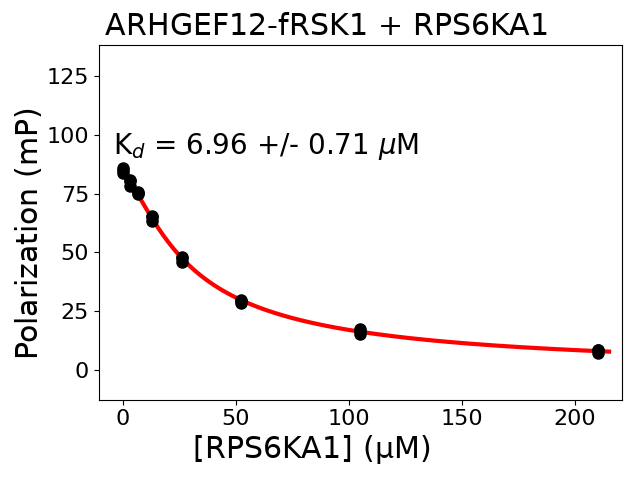
PUBMED ID of publication: 36115835
Number of measurements: 3
Measured pKd: 5.16
Experiment series 4
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (SEP)732
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPpSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6M): 98.42
Standard deviation of dissociation constant (10-6M): 11.17
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit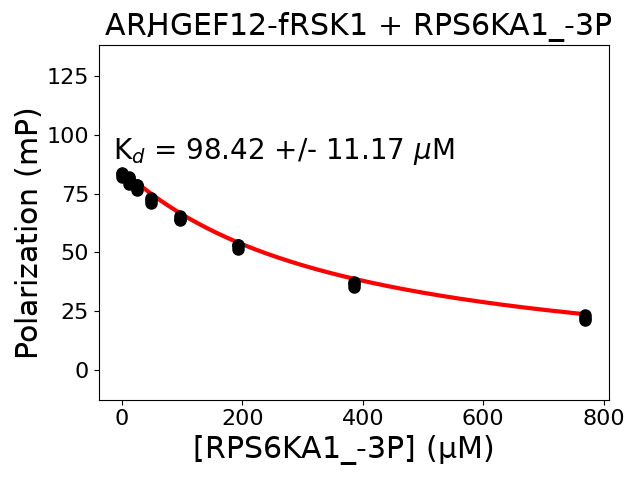
PUBMED ID of publication: 36115835
Number of measurements: 3
Measured pKd: 4.01
Experiment series 5
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (TPO)733
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPSpTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6M): 0
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit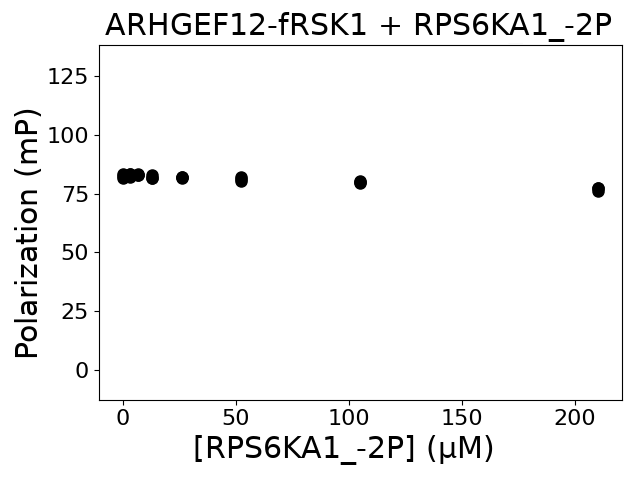
PUBMED ID of publication: 36115835
Number of measurements: 3
The affinity was below the detection threshold of the assay.
Experiment series 6
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (TPO)734
Protein A sequence: RVRKLPSTTL
Protein A construct: Biotin-ttds-RRVRKLPSTpTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6M): 13.26
Standard deviation of dissociation constant (10-6M): 1.47
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit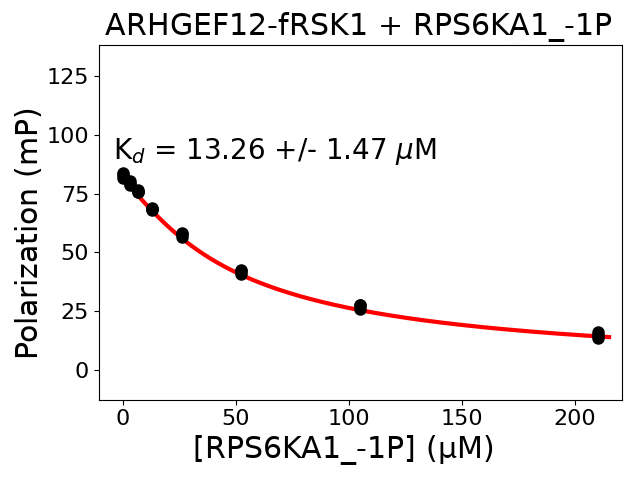
PUBMED ID of publication: 36115835
Number of measurements: 3
Measured pKd: 4.88
Experiment series 7
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (SEP)732
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPpSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Average holdup BI: 0.08
Normalized holdup BI: 0.08
Immobilized partner concentration in holdup experiment (10-6M): 15
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 3.79
Experiment series 8
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (TPO)733
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPSpTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Average holdup BI: 0.04
Normalized holdup BI: 0.05
Immobilized partner concentration in holdup experiment (10-6M): 18
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 9
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (TPO)734
Protein A sequence: RVRKLPSTTL
Protein A construct: Biotin-ttds-RRVRKLPSTpTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Average holdup BI: 0.48
Normalized holdup BI: 0.62
Immobilized partner concentration in holdup experiment (10-6M): 25
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 4.87
Experiment series 10
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Average holdup BI: 0.73
Normalized holdup BI: 0.81
Immobilized partner concentration in holdup experiment (10-6M): 18
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
PUBMED ID of publication: 36115835
Number of measurements: 1
Measured pKd: 5.45
Experiment series 11
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (SEP)732
Protein A sequence: RVRKLPSTTL
Protein A construct: biotin-ttds-RRVRKLPpSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Average holdup BI: 0.11
Normalized holdup BI: 0.14
Immobilized partner concentration in holdup experiment (10-6M): 15
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Experimental details: not reported internal standards of holdup layout #5
Number of measurements: 1
Measured pKd: 4.05
Experiment series 12
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A sequence: RVRKLPSTTL
Protein A construct: Biotin-ttds-RRVRKLPSTTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6M): 2.79
Standard deviation of dissociation constant (10-6M): 0.11
Experiment method: SPR
PUBMED ID of publication: 30726710
Number of measurements: 1
Measured pKd: 5.55
Experiment series 13
Protein A protein: RPS6KA1
Protein A fragment: 725-735
Protein A site: PBM
Protein A modification: (SEP)732
Protein A sequence: RVRKLPSTTL
Protein A construct: Biotin-ttds-RRVRKLP(pS)TTL
Protein B protein: ARHGEF12
Protein B fragment: 67-151
Protein B site: PDZ
Protein B construct: his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6M): 0
Experiment method: SPR
PUBMED ID of publication: 30726710
Number of measurements: 1
The affinity was below the detection threshold of the assay.